
Manually select single worms for worm modeling.Pre-process data by compensating for non-uniform illumination and removing artifacts such as well edges and bubbles.This pipeline requires manual interaction selecting representative non-touching worms. Steps A-C are the same as in Pipeline 1 the same pre-processing and foreground/background segmentation should be applied both when creating the worm model and when running the analysis as changes to the pre-processing may affect the appearance of the width of the worms, in turn influencing the model. DefaultWormModel.xml provided with Pipeline 1) fits the new data. This pipeline may be omitted if a previously created model (e.g. Pipeline 2: Find, select, and save individual wormsĬellProfiler 2.1 pipeline- Pipeline2_SelectSingleWorms.cppipe Export measurements to an SQLite database, save worm segmentation masks and images of worm and fat outlines.Identify sub-regions (fat) and extract measurements.Straighten worms and extract measurements.Identify individual worms by worm untangling and extract measurements.Separate worms and worm clusters the from image background by foreground/background image segmentation.Pre-process data by removing artifacts such as well edges and bubbles.This pipeline is fully automated, does not need any user input (once optimized), and can be run on a large number of images organizing output based on information extracted from the input file names. 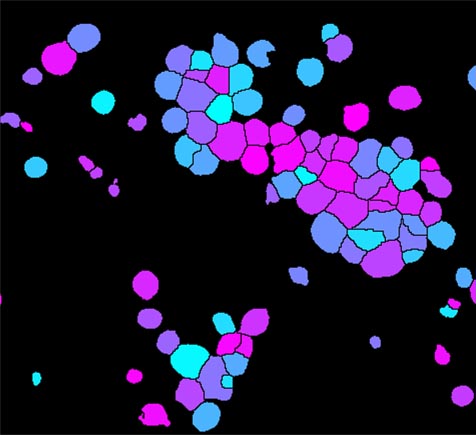
This model could either be the DefaultWormModel.xml provided above, or a new worm model created by pipelines 2 and 3 above. 
This is the main pipeline for delineating worms (also those in clusters) and extracting different kinds of shape and intensity measurements. Pipeline 1: Identify and collect measurements from individual worms and sub-regionsĬellProfiler 2.1 pipeline- Pipeline1_UntangleWormsExtractMeasurements.cppipe Note that CellProfiler 2.1 should be used. All measurements are exported to a database for further exploration and phenotyping. may also be extracted from individual worms after manual correction. Measurements, such as intensity of stain, fat stain distribution, fat region size, worm width etc. due to a different image resolution or different worm strain), a new worm model can be created using Pipeline 2 (pink) to (g) manually select non-touching worms making up a training set for a new model built by Pipeline 3.įor low-throughput experiments, it is possible to manually curate faulty segmentation results using Pipeline 4 (orange) by (h) manual editing of the output from step d of pipeline 1, and (i) manual flipping of digitally straightened worms so that they all align in with heads or tails up. If the provided default worm model does not fit the input data (e.g. Once worms and fatty regions are identified, a large number of intensity, shape and texture measurements are extracted. Now, each individual detected worm has a unique color.ĭigitally straighten worms and align them to a simple worm atlas for measurement of stain distribution.ĭetect fatty regions within worms. Individual worms/worm clusters are randomly colored.ĭelineate individual worms by model-based worm untangling. Pre-process to detect artifacts such as bubbles and well edges.ĭetect worms and clusters of worms. The main analysis pipeline, Pipeline 1, shown in blue, is fully automated, and consists of the following steps:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
